1st Prize winner: Anna Franz, School of Biochemistry, University of Bristol.
 After completing my undergraduate studies in Biology in Heidelberg, I moved to the Gurdon Institute in Cambridge to do my PhD in Prof. Jordan Raff’s lab as part of the Wellcome Trust-funded PhD programme in Developmental Biology, where I studied the role of the centrosomal protein CP110 in centriole duplication in Drosophila. I’m now a postdoctoral researcher in the labs of Prof. Paul Martin and Prof. Will Wood in Bristol studying the role of fat body cells during wound healing in Drosophila.
After completing my undergraduate studies in Biology in Heidelberg, I moved to the Gurdon Institute in Cambridge to do my PhD in Prof. Jordan Raff’s lab as part of the Wellcome Trust-funded PhD programme in Developmental Biology, where I studied the role of the centrosomal protein CP110 in centriole duplication in Drosophila. I’m now a postdoctoral researcher in the labs of Prof. Paul Martin and Prof. Will Wood in Bristol studying the role of fat body cells during wound healing in Drosophila.
The fat body, the fly equivalent of the mammalian liver and adipose tissue, is a versatile tissue, comprised of very large cells, that performs several important systemic functions throughout the fly life cycle. These disparate roles include growth control, storage and regulation of lipid and carbohydrate delivery and the production of antimicrobial peptides after infection. We think that all of these systemic functions of the fat body could be pivotal but required locally at any site of tissue wounding, and are currently investigating this in the Drosophila pupa.
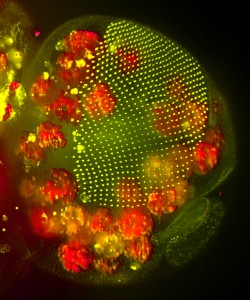
This image, taken with a light sheet microscope, shows the head of a Drosophila pupa: The developing compound eye (green) is composed of several hundred simple units called ommatidia arranged in an extremely regular array. The giant polyploid cells of the fat body (red) occupy a large volume of the head.
2nd Prize Winner: Ronan Mellin, MRC Human Genetics Unit, Edinburgh.
I’m an MRC-funded PhD student in Dr Luke Boulter’s lab at the MRC Human Genetics Unit in Edinburgh. I am looking at the role of non-canonical Wnt signalling in the adult intestine to better understand how this signalling pathway governs tissue architecture in health and disease.
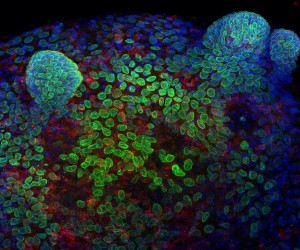
Here, I am using murine colonic epithelial organoids as an ex-vivo tool to model intestinal cell polarity. Colonic crypts were isolated and grown in Matrigel (a 3D matrix). Following culture, projections containing the epithelial stem cells and progenitors can be seen protruding from the spheroid. These represent the colonic crypt, and can be easily manipulated to define the signals that regulate epithelial architecture. This organoid is stained for DNA with DAPI (Blue), the nuclear envelope with LaminB1 (Green) and the centrosome marker γ-tubulin (Red). This method gives us a highly tractable three-dimensional model of regeneration, which we can use to computationally study cellular processes.
3rd Prize Winner: Helen Weavers, School of Biochemistry, University of Bristol.
After graduating with a degree in Developmental Biology, I embarked on a Wellcome Trust funded PhD in Prof. Helen Skaer’s lab on the Developmental Biology 4-Year PhD Programme at the University of Cambridge. Here, I have been studying the development of the fly’s excretory (renal) system, focusing on the role of specialised tip cells that guide the formation of the kidney’s complex 3D architecture which is key to optimal physiological functioning. I am now continuing my interest in developmental biology as a Post-doctoral Research Associate in Biomedical Sciences at the University of Bristol in the labs of Prof. Paul Martin and Prof. Will Wood where I am studying the development and function of the Drosophila immune system.
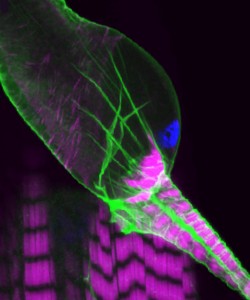
This confocal image shows the intricate structure of the developing fly kidney, which is anchored within the body by attachment to nearby heart muscle. The striking striations of the heart (and body wall muscle beneath) are revealed by labeling Actin (majenta). Cell membranes (green) and nuclei (blue) are also stained.