Written for the BSCB by Dr Ian Cowell, Institute for Cell and Molecular Biosciences, University of Newcastle-Upon-Tyne, UK
Quick look: In its modern sense, epigenetics is the term used to describe inheritance by mechanisms other than through the DNA sequence of genes. It can apply to characteristics passed from a cell to its daughter cells in cell division and to traits of a whole organism. It works through chemical tags added to chromosomes that in effect switch genes on or off.
Researchers studying the microscopic roundworm Caenorhabditis elegans recently discovered a set of mutations that extended the worms’ normal 2-3 week lifespan by up to 30%. This was exciting, not least because discoveries in animals such as roundworms can sometimes help us understand processes like ageing in humans. This was not the end of the story though, as the researchers found that the descendants of the long-lived roundworms could also live longer than normal, even if they only inherited the non-mutated version of the genes from their parents. This doesn’t seem to make sense at first; surely characteristics such as hair colour, height and even how long we or a microscopic worm could potentially live are carried in the DNA sequence of the genes that we inherit from our parents. So how can we solve the conundrum of how the roundworms inherited the long lived characteristic, without inheriting the DNA sequence that initially caused it? The answer is epigenetics.
It’s not all in your DNA
In a nutshell, epigenetics is the study of characteristics or “phenotypes” that do not involve changes to the DNA sequence; and the long-lived roundworms are just one of many examples. Others, as we will see below, include how queen and worker honey bees can appear so different despite being genetically identical, how starvation in human populations may affect the health and longevity of the next generation, why all tortoiseshell cats are female and even how we all develop from a single cell (a fertilized egg) to end up with bodies containing many different types of specialised cells but which all contain the same genes and DNA sequence.
So what is epigenetics?
Another way of looking at epigenetics is like this; while traditional genetics describes the way the DNA sequences in our genes are passed from one generation to the next, epigenetics describes passing on the way the genes are used. To make a computer analogy, think of epigenetics as metadata, information describing and ordering the underlying data. If you own an MP3 player for example, it will contain a lot of data, the MP3 files. Think of these as analogous to genes. But you will also probably have playlists or you may play tracks by artist or genre. This information, playlist, artist, genre etc. is metadata. It determines which tracks are played and in what order, and this is what epigenetics is to genetics. It is a set of processes that effects which genes are switched-on, or “expressed”, as molecular biologists would say.
How does epigenetics work?
So epigenetics is about how genes are expressed and used, rather than the DNA sequence of the genes themselves, but how does this work? Many researchers have been studying epigenetics over the past few decades, and it is currently an area of intense research activity. We know that a part of how epigenetics works is by adding and removing small chemical tags to DNA. You can think of these tags as post-it notes that highlight particular genes with information about whether they should be switched on or off. In fact the chemical tag in question is called a methyl group (see Diagram 1) and it is used to modify one of the four bases or “chemical letters”, A, C, T and G, that makes up the genetic code of our DNA. The letter that is tagged is C or cytosine and when it is modified, or methylated it is called 5-methyl cytosine. Methyl groups are added to DNA by enzymes called DNA methyl transferases (DNMTs).
-
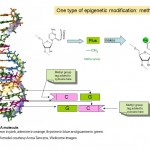
Diagram 1. Two chemical tags, methyl and acetyl groups that are central to epigenetic phenomena and the chemical structure of cytosine and 5-methyl cytosine in DNA. The pentagonal part of the molecule forms the continuous “backbone” of the DNA . Only one of the two strands of DNA that makes up the familiar double helix is shown.
Queen bee status is partly determined by fewer methyl tags
In most cases, more methylated Cs in the DNA of a gene results in the gene being switched off. Honey bees provide us with a good example of how this can work. Worker bees and the queen have very different bodies; the queen is much larger, longer lived, has an enlarged abdomen and lays many thousands of eggs, while the smaller workers are sterile but have complex foraging and communication skills. Despite this, the queen and workers in a hive are female and genetically identical. The clue to how this comes about lies in royal jelly, a secretion that is fed to some developing larvae, and which results in these larvae becoming queens rather than workers. We will come back to royal jelly and its queen-making properties later, but a fascinating piece of research showed that if the amount of the methyl group adding DNMT enzyme was artificially reduced in bee larvae, then the larvae developed into queens, even if they weren’t fed royal jelly. Thus, the switch between queen and worker can be flipped by the abundance of methyl tags on the bee larvae’s DNA. Fewer methyl tags leads to switching on of a special gene or genes in the developing larvae that results in the development of the larvae into queens and not workers.
Tags on tails also operate gene switches
DNA methyl tags are only one part of the story though. In the cells of all plants and animals, DNA is packaged or wrapped up into nucleosomes where the DNA double helix is wrapped around a central core of protein (see Diagram 2). About 150 letters-worth of DNA (or base-pairs) is wrapped around each nucleosome, and this helps package the 3 billion base pairs of genetic code into each of our cells. Nucleosomes are too small to see using conventional microscopes, but biologists use a technique called X-ray diffraction to work out the shape and organisation of objects like nucleosomes, and in 1997 this technique revealed the beautiful structure of nucleosomes at high resolution – see (http://www.rcsb.org/pdb/explore/explore.do?structureId=1aoi).
-
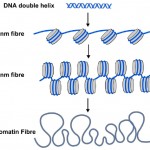
Diagram 2. The familiar DNA double helix (blue) is wrapped around nucleosomes (grey cylinders) in cells. The string of nucleosomes can be coiled into a thicker filament, called the 30 nm fibre and this can be further coiled into a still thicker chromatin fibre. When genes are switched on their nucleosomes are more uncoiled like the 10nm fibre.
Nucleosomes are compact, but the ends or “tails” of the proteins that make up the nucleosome, which are called histones, stick out from the otherwise compact nucleosome structure. Like the methyl tags on DNA, small chemical tags can also be added to these histone tails (see Diagram 3). Two of the chemical tags that are added to these tails are acetyl groups and methyl groups. Methyl, acetyl and a few other types of tags can be added to the tails in a large number of combinations and this effects whether an underlying gene is switched on or off. In fact genes can be switched right off (this is called silencing), full on, or somewhere in between by DNA methyl tags and histone tail tags. The combination of DNA and histone tags can also effect how easily a gene is turned on or off.
When cells divide
When cells divide, the entire DNA sequence from the original cell (3 billion base pairs contained in 23 pairs of chromosomes in a human cell) is duplicated so that both daughter cells receive an exact copy. What, you might ask, happens to all those epigenetic tags? We have known for some time that the DNA-methyl tags are copied too, so that both daughter cells have the same pattern of DNA methylation. We now know that the pattern of histone tags is also mostly duplicated as cells divide, although this is currently less well understood. Nevertheless, cell division is also a time when epigenetic tags can most easily be changed.
Return of the long-lived worm
Right at the beginning we came across the story of the long-lived microscopic worms thatpassed on their longevity to their offspring even if the individual offspring did not inherit the variant gene (mutation) that originally caused the extended lifespan. We are now in a position to explain this apparently strange result. In most cases genes contain the information to make a protein molecule, and the protein molecules might be enzymes that carry out chemical reactions in the cell, or parts of the structure of the cell itself. It turns out that the genes that were mutated in the worm study make proteins that work together to add a methyl tag to nucleosomes. This tag is an on-switch. When one or more of the genes were mutated this tag was absent and several genes that should be on, including some involved in ageing, were switched off and the worms had a longer lifespan. The unexpected thing is that the epigenetic tags were thought to be completely erased or reset during the formation of sperm and egg, and so unlike the genes themselves they shouldn’t be passed on to the next generation. But this result and other research that shows that this is not always the case and that sometimes, the pattern of epigenetic tags are passed on.
How to make a queen
Whether a larval honey bee becomes a worker or a queen depends on an epigenetic switch, and this switch seems to be “flipped” by royal jelly. But what is it about royal jelly that leads a larva that would otherwise grow up to be a worker, to become a queen? The answer lies in understanding that the individual chemical tags that are added to the histone tails of nucleosomes are constantly being revised by the cell. Acetyl tags are added by enzymes called histone acetyl transferases and they are removed or erased by a second group of enzymes called histone deacetylases (HDACs). Both of these enzymes are present in most cells and this allows genes to be switched on or off over time.
More acetyl tags help deliver queen bee status
Recently, researchers set out to identify compounds in royal jelly that could alter this process, and what they found was something known as an HDAC inhibitor. This was a relatively simple chemical compound that is present in royal jelly and that stops the action of HDAC enzymes that normally remove acetyl tags from histones. This results in a build-up of acetyl tags in the cells of the bee embryos, and like the reduction in DNA-methyl groups described previously, this is thought to switch on key genes required for development of a queen. Without the HDAC inhibitor in the royal jelly, the larvae follow a “default” set of genetic instructions and develop into workers.
HDAC inhibitors are not only important to queen bees, but are also part of a small but growing number of medically useful drugs that target epigenetic tags and which are useful in treating some kinds of cancer. Furthermore HDACs also have a role in the way our brains form memories, and novel drugs that affect histone acetylation may have a role in the future in treating memory impairment in elderly patients.
The environment and epigenetics
We have seen how the difference between a queen and worker bee is determined by exposure to a chemical that directly alters epigenetic tags such as acetyl groups; but are there examples where nutrition or other aspects of the environment affect human populations in a way that can be explained by epigenetics? Obviously we can’t do experiments on human populations as we can on microscopic worms or bees, but sometimes human history or natural phenomena do it for us. One such example is what is known as the Dutch Hunger Winter. In the last year of the Second World War in Europe, a food embargo imposed by occupying German forces on the civilian population of the Netherlands resulted in a severe famine, coinciding with a particularly harsh winter. About 20,000 people died from starvation as rations dropped to below 1000 kilocalories per day. Despite the chaos of war, medical care and records remained intact allowing scientists to subsequently study the effect of famine on human health. What they found was that children who were in the womb during the famine experienced a life-long increase in their chances of developing various health problems compared to children conceived after the famine. The most sensitive period for this effect was the first few months of pregnancy. Thus, something appears to happen early in development in the womb that can affect the individual for the rest of their lives.
Epigenetic effects can sometimes pass to grandchildren
Even more surprisingly, some data seems to suggest that grandchildren of women who were pregnant during the Hunger Winter experience some of these effects. From what we have already discussed, this strongly suggests an epigenetic mechanism. In fact, research with the Dutch Hunger Winter families continues, and a recent study looking at a gene galled IGF2 found lower levels of the methyl tag in the DNA of this gene in individuals exposed to the famine before birth. Although IGF2 may not itself be involved in the increased risk of poor health in these people, it shows that epigenetic effects (i.e. reduction of the number of methyl tags on particular genes) that are produced before birth can last for many decades. Studies in animals have also found that the diet of the mother can have effects on her offspring. For example, feeding sheep a diet lacking the types of food required to make methyl groups leads to offspring with altered patterns of DNA methylation and which have higher than expected rates of certain health problems.
Epigenetics and imprinting, why genes from Mum and Dad are not always equivalent
We all have 23 pairs of chromosomes in our cells. For each pair, one came from mother and one from father. Thus, we inherit one copy of each gene from each parent and we generally assume that the function of the gene does not to depend on which parent it came from. However, for imprinted genes things are different. For these genes, either the maternal or paternal copy of the gene is active, while the other one is kept silent. There are at least 80 imprinted genes in humans and mice, many of which are involved in growth of the embryo or the placenta. How can one copy of a gene be switched off, while the other copy in the same cell is switched on? The answer is epigenetics. Probably the most studied imprinted gene is IGF2(see above). One part of IGF2 operates as a switch. If the DNA is methylated here the IGF2 gene can be expressed. The switch is only methylated in Dad’s copy of the gene and so only this copy is expressed, while the maternal copy is silent. This switch is thought to be set up in the gametes (eggs and sperm) so right from the start, genes received from Mum and those from Dad are labelled differently with epigenetic tags and so are not equivalent.
Imprinting and mental disorders
Angelmann and Prader-Willi syndromes are two distinct genetic conditions with different symptoms, both caused by loss of a part of chromosome 15. Children who inherit one copy of this faulty chromosome develop either Angelmann or Prader-Willi syndrome, despite having a normal copy of the chromosome from their other parent. So how does the same mutation (loss of part of chromosome 15) lead to these two different conditions? The answer lies in the discovery that this particular piece of chromosome 15 contains a number of genes that are imprinted, so only the paternal or maternal copy of these gene are expressed; which of the two syndromes appears depends on whether the deletion was in the maternal or paternally inherited chromosome. When the faulty chromosome is inherited from Dad, there is no functional copy of the imprinted genes that are switched off on the maternal chromosome 15 and the result is Angelmann syndrome and vice versa for Prader-Willi syndrome. This is quite unlike most genetic conditions such as cystic fibrosis, where an effect on development or health is only seen when a mutated gene or genes is inherited from both parents.
Boys versus Girls, how to switch off a whole chromosome
A bit of genetics that most of us know about is what makes a boy a boy, and a girl a girl. It’s the X and Y chromosomes. At the very beginning of our existence each of us received one X chromosome from our Mums via the egg, and while the girls received another X chromosome from their dads, via the sperm, the boys got a Y chromosome. The Y chromosome in the cells of a male embryo directs it to develop into a boy, while with two X and no Y chromosome the female embryo develops into a girl. Now, you might notice that there is an imbalance here. We all have two each of all the other chromosomes, but for the sex chromosomes (X and Y) the girls have two Xs while the boys only have one X (and a Y). While the Y chromosome contains few genes, mostly involved in “maleness”, the X chromosome contains quite a few genes involved in important processes such as colour vision, blood clotting and muscle function. In order to even up the “dosage” of X chromosome genes between male and female cells, one entire X chromosome is switched off in female cells. This is called X-chromosome inactivation and happens very early in the womb. In this process cells randomly switch off either the paternal or maternal X chromosome, so that when a girl baby is born her body is a mixture or chimera of cells where either the maternal or paternal X-chromosome is switched off. The way that this happens involves the type of epigenetic tags that we have discussed and it has been known for decades that female cells contain one very compact X chromosome called the Barr body that can be seen under the microscope, and this is the inactive X chromosome.
The case of the tortoiseshell cat
We are probably all familiar with tortoiseshell cats and their mottled coats with patches of orange and black fur. What you might not know is that almost all cats with this type of coat are female! The reason for this is that a gene for coat colour is located on the cat’s X chromosome. There are two versions of this gene, called “O” and “o”; one gives ginger fur and the other black. Two copies of the same version in a female cat results in ginger or black fur respectively, but one copy of each gives a tortoiseshell effect. This is down to X-chromosome inactivation. The skin of these cats is composed of patches of cells where either the maternal or paternal X chromosome is inactivated. This results in skin with the O gene switched on and o silenced in some patches (orange fur) and o gene on and O silenced in other patches (black fur), hence the tortoiseshell pattern. Since the male cats only have one X chromosome, and no X-chromosome inactivation, they are either orange or black all over.
Epigenetic inheritance, can epigenetic states be passed from one generation to the next?
As we have seen from the roundworm example, epigenetic effects (in this case extended lifespan) can sometimes be passed from one generation to the next, although the effects only seem to last for a few generations. Are there examples where epigenetic effects carry over to subsequent generations in humans or other mammals? There is some evidence that the effects of the Dutch Hunger Winter affected grandchildren of women who were pregnant during the famine. Similarly, in a study of a 19th century northern Swedish population who underwent cycles of famine and plenty, the amount of food available appears to have affected the health and longevity of the next generation.
Hair colour in mouse can be determined by an epigenetic effect
Perhaps the best known example of transgenerational epigenetic effects is provided by the mouse Agouti gene. This gene controls hair colour, and is switched on at just the right time in hair follicle cells to produce a yellow stripe in the otherwise dark hairs, resulting in what is called an agouti coat. But mice with a particular variant of the Agouti gene called Avy have coats that are anywhere between yellow and the normal dark (agouti) pattern of wild-type mice. The yellow mice also become obese and suffer other health problems. So the Avy gene seems to have a variable effect (in fact the Avy stands for Avariable yellow). How this works has puzzled geneticists for years, but we can now recognise this as an epigenetic effect. The yellow fur occurs because Avy version of the Agouti gene has faulty controls and is switched on all the time. However, methyl tags are often added to the faulty control DNA sequence and this tends to switch the gene off, resulting in mottled or dark agouti fur in individual mice. Pups born to dams with the Avy gene range in colour from yellow to dark, but the proportion depends on the coat colour of the mother; litters of dark (agouti) females are more likely to contain dark pups. Furthermore, a higher proportion of dark offspring is observed if both the mother and the grandmother have the dark colouration. So the agouti colouration, which is determined epigenetically (by the number of methyl tags on the Avy gene) can to some extent, carry through from one generation to the next.
Eggs and sperm do not usually ‘carry over’ epigenetic effects
Although we can find cases where epigenetic effects apparently last from parents to offspring, this is not usually the case and almost all of the epigenetic switches or marks are reset in germ cells (eggs and sperm) and in the very earliest stages of development of an embryo. In fact if this wasn’t the case, the amazing development of a fertilised egg into a fully formed creature would be impossible.
Getting from a fertilized egg to a fully formed human, it’s all in the (epi) genome
So far we have described some specific cases of epigenetic regulation, but we now know that epigenetics in its broad sense, (how genes are expressed and used, rather than the DNA sequence of the genes themselves) is central to how a fertilised egg can eventually give rise to a whole organism and how cells of, let’s say your skin, remain skin cells and are different from your brain cells, despite containing exactly the same genes. Shortly after fertilisation, a developing human embryo consists of a ball of cells called embryonic stem cells. Each of these cells has the capacity to give rise to any of the types of cells in the body as the embryo grows (for example, brain cells, skin cells or blood cells). By contrast, 9 months later when a baby is born, most of the cells making up his or her body are committed to be a specific type of cell with specific functions. So as the cells divide, the ball of embryonic stem cells gradually develops into all the cell types and structures of the baby at term. For this to happen, thousands of genes must be switched on or off at just the right times and in the right cells as an embryo grows. For example, genes that make the fibrous keratin protein that gives our skin its strength, are only switched on in skin cells and not in the developing brain and genes required for brain cells to develop and make their interconnections are on in the brain but not in the skin.
During development genes have to be switched ‘on’ and ‘off’. Epigenetic tags help with this
A very big area of research today concerns how all this gene switching on an off works, and a large part of this process uses the epigenetic chemical tags, especially acetyl and methyl histone tags. In order for those embryonic stem cells to be able to give rise to all of the other types of cells, their epigenetic switches are (almost) completely reset compared to adult cells. I have put “almost” in brackets as we know from imprinted genes and transgenerational epigenetic inheritance that there are exceptions.
Epigenetics, Dolly the sheep and friends
In February 1997, a sheep called Dolly became the most famous example of her species, briefly even becoming a TV celebrity. The reason for her fame is that she was the first mammal to be “created” by a process called somatic cell nuclear transfer, or in other words the first man-made clone (man-made to be distinct from identical twins, who are natural clones). The process leading to her birth required a mature oocyte (a unfertilised egg) from one female sheep and an ordinary cell from the udder of a second sheep. First the nucleus (the part containing the DNA) was removed from the oocyte. This was done using a special microscope as although oocytes are quite big compared to other cells, they are still too small to see with the naked eye. Then the nucleus from the udder cell was inserted into enucleated oocyte. Thus, Dolly had three “mothers”: the donor of the oocyte, the donor of the udder cell and the sheep that carried the developing embryo to term. No father was involved. Although this process was, and remains, very inefficient it was the first proof that the genes from an adult mammalian cell can be “epigenetically reprogrammed” back to the state of the embryonic stem cells that can develop into any other type of cell. Subsequently the same process has been applied to other species and may have medical uses in generating cells that could repair tissues damaged by injury or disease.
Summary: the epigenome and the ENCODE project – the “Large Hadron Collider” of Biology
Whereas the term “genome” refers to the entire DNA sequence of an organism (three billion letters of it for humans), the epigenome refers to the entire pattern of epigenetic modifications across all genes, including methyl DNA tags, methyl histone tags, acetyl histone tags and other chemical tags that we have not mentioned, in each cell type of an organism. This represents an almost unimaginable amount of information, dwarfing even the human genome project. Nevertheless, knowledge of the epigenome is essential to fully answer some of the biggest questions in biology such as: how do we develop from a ball of identical cells into a whole organism? why do we age? and how can we better understand diseases such as cancer? Not surprisingly then, epigenetics and the epigenome is a big area of research. Some of the research in this field is encompassed by the ENCODE (Encyclopedia of DNA Elements) project, an ongoing venture to identify patterns of epigenetic tags in many different types of cells for the entire human genome (http://genome.ucsc.edu/ENCODE/). The ENCODE project is sometimes likened to the Large Hadron Collider or LHC in Switzerland. The LHC is the largest piece of scientific equipment ever built and the experiments physicist conduct with it aim to probe the fundamental details of the matter that makes up our Universe. Although biologists don’t have (or need) such a spectacular piece of kit for their research, the effort to examine the intricacy of the human epigenome has been likened to the LHC project because of its scale, complexity and the amount of information being created.
Epigenetic errors
Epigenetics is an area where our scientific knowledge is rapidly increasing. One thing that scientists have discovered is that epigenetic errors are common in diseases such as cancer and in ageing cells. As a result, scientists are developing medicines that target faulty epigenomes and one of the first examples is the use of HDAC inhibitors, similar to the compound found in royal jelly. From the study of strange patterns of inheritance such as genetic imprinting, the yellow/agouti Avy mouse, the all-female tortoiseshell cat population and other related phenomena biologists have uncovered a whole new layer of information that lies “on top” of the DNA sequence of our genes. These new discoveries explain these previous puzzling observations, but also have great potential for new understanding and treatments for human disease.
Further Reading:
- Bird, Adrian. ’Epigenetics. Instant Expert No. 29’, New Scientist, 5th January 2013, No. 2898.
- Carey, Nessa. ‘The Epigenetics Revolution: How Modern Biology is Rewriting Our Understanding of Genetics, Disease and Inheritance’. Publisher: Icon Books. Paperback 1st March 2012. ISBN-10: 1848313470. RRP Price £9-99.
Epigenetic Influences and Disease at Website:
- http://www.nature.com/scitable/tpicpage/epigenetic-influences-and-disease-895 [Accessed 19-11-12]